Association Of Neurotransmitter System Genes With Facial Emotion Recognition And Emotional Intelligence
Abstract
The present work was aimed at studying the facial emotion recognition and emotional intelligence in carriers of different genotypes of BDNF, COMT, HTR2 genes. These genes are associated with neurotransmitters systems and, therefore, can be associated with studied characteristics. To study facial emotion recognition, we recorded hit rates and verbal reaction times in response to video clips depicting facial expressions of six basic emotions (anger, disgust, fear, happiness, sadness, and surprise). Emotional Intelligence by Lyusin, The Emotional Intelligence Self-Evaluation by Hall, and Mayer-Salovey-Caruso Emotional Intelligence Test were used as measures of emotional intelligence. DNA was extracted from buccal cells, genotyping procedure was implemented by PCR method (“Biological Solutions and Technologies”, Russia, Moscow). Analysis of variance with repeated measures showed that carriers of Val/Met of BDNF were more accurate than carriers of Val/Val of BDNF at recognition of anger. In regard to emotional intelligence, carriers of Val/Met of BDNF showed a high level of the general score of Mayer-Salovey-Caruso Emotional Intelligence Test than carriers of Val/Val of BDNF. We did not find any significant effect of genotypes of COMT and HTR2A on facial emotion recognition. Also we did not find any significant effect of genotypes of COMT and HTR2A on emotional intelligence measures.
Keywords: Facial emotion recognitionemotional recognitionBDNFCOMTHTR2Aemotion
Introduction
Emotional intelligence is a phenomenon that has drawn attention in the last years. To date, the most widespread studies of the following aspects of the recognition of emotions and emotional intelligence include: gender differences (Olderbak S. et al., 2018, Wingenbach et al, 2018; Wright, et al., 2018), mental diseases (parkinsonism, autism, schizophrenia, depression; Berggren et al., 2018, Glenthøj et al., 2018, Wasser et al., 2018), the problem of teaching neural networks to the recognition of emotional expression is widely studied (Ko, 2018; Tautkute, 2018). However, the question of the influence of different genotypes of the genes of neurotransmitter systems on the success of the recognition of emotions and the level of emotional intelligence of healthy people is, in our opinion, insufficiently studied.
Catechol-O-methyl transferase gene (COMT) and serotonin receptor 2A gene (HTR2A) attracted our attention due to their connection with the neurotransmitter systems (catecholamines), which, in turn, affect the emotional sphere of the person (Alfimova et al., 2009; Golimbet et al., 2004). BDNF gene controls Brain Derived Neurotrophic Factor, which provides the processes of synaptic transmission, neuroplasticity, and adequate adaptation of the nervous system to changing environmental conditions (Egan et al., 2003; Popova et al., 2017). These genes are associated with the serotonergic system of the brain, namely with the concentration, the time that the neurotransmitter stays in the synaptic cleft, and the sensitivity of serotonin receptors. This allows us to assume the differences between genotypes of the neurotransmitter system genes in the level of emotional intelligence and the accuracy of emotion recognition.
Problem Statement
There are three genotypes on polymorphic loci Val158Met of the COMT gene. The Met / Met genotype is associated with the long stay of catecholamines in the synaptic cleft, with a tendency to various mental illnesses, high anxiety and a tendency to autodestructive behavior; Val / Val - with a short stay of catecholamines in the synaptic cleft and a more harmonious personality profile, the heterozygous Val / Met genotype - with the average length of stay of catecholamines in the synaptic cleft (Opmeer, et al., 2013).
The BDNF gene also has the genotypes Val / Val, Val / Met and Met / Met. The presence of the Met allele is associated with a decrease in the secretion of the cerebral neurotrophic factor. In this regard, carriers of the Met allele have a lower level of adaptive capacity, a higher risk of developing depression and anxiety (Popova et al., 2017).
A special attention of researchers is attracted to 2 mutations of HTR2A gene: 4692G>A, rs6311 (Tr2) и 6230С>T, rs6313 (Tr3). According to modern ideas, alleles A and T are associated with increased expression of serotonin (Polesskaya, 2004). The genotypes G / G and C / C are associated with a high level of social introversion (Golimbet et al., 2004), neuroticism and low level of self-esteem, propensity to depression (Chee et al., 2001; Choi et al., 2004). According to Barskij V. et al. (Barskij et al., 2010), girls-carriers of the G / G genotype are less sensitive to socially acceptable rules, to the people approval or condemnation. With genotypes A / A, T / T associated normative behavior, resoluteness, the pursuit of success, conscientiousness, social extraversion, low level of neuroticism and a higher level of aggression (Golimbet et al., 2004; Chee et al., 2001; Choi et al., 2004; Fiocco et al., 2007).
When sad faces were presented in groups of patients with bipolar, depressive and anxiety disorders, the following features were noted irrespective of the diagnosis: in carriers of the Val allele of the COMT gene the amygdala was activated; in the carriers of the Met allele, hyperactivation of the ventral prefrontal cortex was detected (Lelli-Chiesa et al., 2011). Healthy carriers of the Val allele recognize neutral stimuli less accurately (according to brain activation data); healthy carriers of the Met allele, encoding the unstable form of the enzyme, recognize affective stimuli less accurately (Alfimova et al., 2015).
Research Questions
Three questions were put forward in our study:
If facial emotion recognition, in general, depends on the genotypes of BDNF, COMT, HTR2A.
If recognition of certain basic emotions depends on the genotypes of BDNF, COMT, HTR2A.
If emotional intelligence depends on the genotypes of BDNF, COMT, HTR2A.
Purpose of the Study
The present work was aimed, therefore, at studying the facial emotion recognition and emotional intelligence in carriers of different genotypes of BDNF, COMT, HTR2A genes.
Research Methods
A sample of 49 student volunteers, aged between 19 and 30 (Mean = 21.0; SD = 2.6), participated in the study. All subjects gave informed consent and received course credits for participation in the experiment. All procedures were conducted in accordance with the Declaration of Helsinki.
Verbal reaction time of facial emotion recognition
We employed the paradigm developed and published by us earlier (Kosonogov & Titova, in press). We used 48 video clips that we made from 56 coloured photographs of faces. There were coloured full-face photographs of 8 actors depicting 7 facial expressions: anger, disgust, fear, happiness, sadness, surprise, and neutral. With the help of the software Sqirlz Morph 2.1 by Xiberpix, we made 48 “neutral-emotion” morph pairs (6 emotional expressions × 8 actors). In the photos, we indicated key points of the face for the software to prepare a gradual change of 12 frames from the first neutral expression to the final emotional one. Thus, each video clip contained 12 frames, lasted 962 ms (Figure

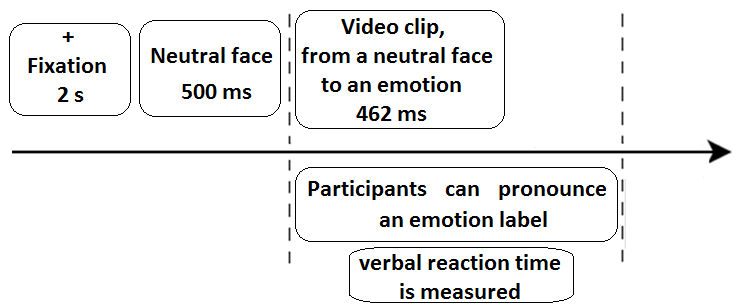
The participants were seated in the experimental chamber in front of a 19-inch computer screen at the distance of 1 m. A microphone (frequency band: 20-16000 Hz; sensitivity: 54 dB; impedance: 2.2 kOhm) was attached to their auricles so that its sensor was 2 cm from their lips and could not be moved by involuntary movements. Participants were told that they would watch video clips depicting 6 basic emotions. The experiment was conducted in OpenSesame (Mathôt, Schreij &Theeuwes, 2012). Each trial began with a fixation period (a cross in the middle of the screen for 2 s), then participants were presented with a video clip. They were asked to announce loudly the emotion label as soon as possible. A technician wrote down participants’ responses (Figure
Emotional intelligence
In order to measure emotional intelligence, we employed Emotional Intelligence by Lyusin (Lyusin, 2009), The Emotional Intelligence Self-Evaluation by Hall, adapted for Russian samples by Fetiskin, Kozlov, Manuylov (Fetiskin, Kozlov, & Manuylov, 2002), and Mayer-Salovey-Caruso Emotional Intelligence Test (MSCEIT; Mayer, Salovey, Caruso, & Sitarenios, 2003), adapted for Russian samples by Sergienko and Vetrova (Sergienko &Vetrova, 2010).
Genetic analysis
DNA was extracted from buccal cells. The genotyping procedure was carried out by using PCR method (“Biological Solutions and Technologies”, Russia, Moscow).
The following DNA sections were analyzed: BDNF gene (68690G>A, rs6265, Val66Met; possible genotypes: Val/Val, Val/Met, Met/Met), COMT gene (23753G>A, rs4680, Val158Met; possible genotypes: Val/Val, Val/Met, Met/Met), HTR2A gene (4692G>A, rs6311 (Tr2), possible genotypes: G/G, G/A, A/A; 6230С>T, rs6313 (Tr3), possible genotypes: C/C, T/C, T/T).
Data analysis
Each dependent variable (hit rates, reaction times, three emotional intelligence measures) was analysed by three independent analyses of variance with repeated measures (one for each gene): Genotype (Val/Val and Val/Met for BDNF and COMT, and G/G, G/A, and A/A for HTR2A × Emotion (anger, disgust, fear, happiness, sadness, surprise); Emotion always being a within-subjects variable. For the main statistical tests, a measure of the effect size, partial eta-squared (
Findings
Facial emotion recognition
We found a marginally significant main effect of genotype of BDNF on hit rates,
We did not find any significant effect of genotypes of COMT and HTR2A on facial emotion recognition (all
Emotional Intelligence
We found a significant main effect of the genotype of BDNF only on the general MSCEIT score,
Conclusion
7.1. Carriers of Val/Met genotype of BDNF showed a better accuracy of facial emotion recognition, provided only by a better recognition of anger. This can be explained by the fact that the BDNF Met gene allele is associated with a high level of anxiety, which provides the most careful analysis of threatening stimuli.
7.2. We did not find any effect of the genotype of BDNF, COMT and HTR2A on reaction time of facial emotion recognition. We can suppose that individual differences in reaction time of facial emotion recognition are provided by other genes, but not the genes under study.
7.3. Curiously, we revealed that carriers of Val/Met genotype of BDNF also showed a higher level of emotional intelligence, as measured by Mayer-Salovey-Caruso Emotional Intelligence Test. This fact indirectly proves that facial emotion recognition can be considered a part of emotional intelligence.
7.4. Two self-report measures, Emotional Intelligence by Lyusin and The Emotional Intelligence Self-Evaluation by Hall, did not show any relationship to BDNF, COMT and HTR2A genes and cannot be recommended in future research upon these genes.
Acknowledgments
The present work was supported by Russian Fund for Basic Research (RFBR) № 18-013-01019.
References
- Alfimova, M., Golimbet, V., Barkhatova, A., Golubev, S., & Korovaitseva, G. (2009). Rol' genotip-sredovyh vzaimodejstvij v razvitii simptomov trevogi i depressii pri stresse, svjazannom s bolezn'ju chlena sem'i. [The role of genotype-environment interactions in the development of symptoms of anxiety and depression related to the disease burden for family] Zhurnal Nevrologii i Psihiatrii im. S.S. Korsakova, 12, 50-54 (in Russian).
- Alfimova, M.V., Mel'nikova, T.S., & Golimbet, V.E. (2015). Molekuljarno-geneticheskie i jelektrojencefalograficheskie markery kognitivnyh processov pri depressivnyh rasstrojstvah [Molecular-genetic and electroencephalographic markers of neurocognitive processes in depressive disorders]. Zhurnal Nevrologii i Psihiatrii im. S.S. Korsakova, 115, 103-109 (in Russian).
- Barskij, V., Aksenova, M.G., Kozlova, O.B., Kirillov, A.V., Demin, A.A., Il'inyh, L.M., & Asanov, A. Ju. (2010). Analiz associacij polimorfnyh markerov genov dofaminergicheskoj (DRD2/ANKK1) i serotoninergicheskoj (HTR2A) sistem mozga s lichnostnymi harakteristikami podrostkov [An association study of polymorphisms of the genes involved in the dopaminergic (DRD2/ANKK1) and serotonergic (HTR2A) brain systems with personality characteristics of adolescents]. Ecological Genetics, 8, 9-17 (in Russian).
- Berggren, S., Fletcher-Watson, S., Milenkovic, N., Marschik, P. B., Bölte, S., & Jonsson, U. (2018). Emotion recognition training in autism spectrum disorder: A systematic review of challenges related to generalizability. Developmental Neurorehabilitation, 21, 141-154.
- Chee, I.S., Lee, S.W., Kim, J.L. (2001). 5-HT2A receptor gene promoter polymorphism-1438A/G аnd bipolar disorder. Psychiatric Genetic, 11, 111-114.
- Choi, M. J., Lee, H.J., Lee, H.J. (2004). Association between major depressive disorder and the -1438A/G polymorphism оf the serotonin 2A receptor gene. Neuropsychobiology, 49, 38-41.
- Egan, M.F., Kojima, M., Callicott, J.H., Goldberg, T.E., Kolachana, B.S., Bertolino, A., Zaitsev, E., Gold B., Goldman, D., Dean, M., Lu, B., Weinberger, D.R. (2003). The BDNF val66met polymorphism affects activity-dependent secretion of BDNF and human memory and hippocampal function. Cell, 112, 257-269.
- Fetiskin, N.P., Kozlov, V.V., & Manuylov, G.M. (2002). Social'no-psihologicheskaya diagnostika razvitiya lichnosti i malyh grupp [Social psychological diagnostics of personality development and small groups]. Moscow: Izd-vo Instituta Psihoterapii (in Russian).
- Fiocco, A.J., Joober, R., Judes, P., Sonia, L. (2007). Polymorphism of the 5-HT2A receptor gene: Association with stress-related indices in healthy middle-aged adults. Frontiers in Behavioral Neuroscience, 1, 3.
- Glenthøj, L.B., Fagerlund, B., Bak, N., Hjorthøj, C., Gregersen, M., Kristensen, T.D., & Nordentoft, M. (2018). Examining speed of processing of facial emotion recognition in individuals at ultra-high risk for psychosis: Associations with symptoms and cognition. Schizophrenia Research, 195, 562-563.
- Golimbet V.E., Alfimova M.V., Mitjushina N.G. (2004). Polimorfizm gena receptora serotonina (HTR2A) i osobennosti lichnosti [Polymorphism of serotonin receptor gene (HTR2A) and personality properties]. Molekuljarnaja Biologija, 38, 404-412 (in Russian).
- Ko, B. C. (2018). A Brief Review of Facial Emotion Recognition Based on Visual Information. Sensors (Basel), 18, E401.
- Kosonogov V. & Titova A. (in press). Recognition of all basic emotions varies in accuracy and reaction time: a new verbal method of measurement. International Journal of Psychology.
- Lelli-Chiesa, G., Kempton, M.J., Jogia, J., Tatarelli, R., Girardi, P., Powell, J., Collier, D.A., Frangou, S. (2011). The impact of the Val158Met catechol-O-methyltransferase genotype on neural correlates of sad facial affect processing in patients with bipolar disorder and their relatives. Psychological Medicine, 41, 779-788.
- Lyusin, D. V. (2009). Oprosnik na ehmocional'nyj intellekt EmIn: novye psihometricheskie dannye [Emotional intelligence questionnaire, EmIn: new psychometric data]. In: Social'nyj i Emocional'nyj Intellekt: ot Modelej k Izmereniyam [Social and Emotional Intelligence: from Models to Measurement], pp. 264-278 (in Russian).
- Mathôt, S., Schreij, D., & Theeuwes, J. (2012). OpenSesame: An open-source, graphical experiment builder for the social sciences. Behavior Research Methods, 44, 314-324.
- Mayer, J. D., Salovey, P., Caruso, D. R., & Sitarenios, G. (2003). Measuring emotional intelligence with the MSCEIT v. 2.0. Emotion, 3, 97-105.
- Molendijk, M.L., van Tol, M.J., Penninx, B.W., van der Wee, N.J., Aleman, A., Veltman, D.J., Spinhoven, P., Elzinga, B.M. (2012). BDNF val66met affects hippocampal volume and emotion-related hippocampal memory activity. Translational Psychiatry, 2, e74.
- Olderbak, S., Wilhelm, O., Hildebrandt, A., & Quoidbach, J. (in press). Sex differences in facial emotion perception ability across the lifespan. Cognition and Emotion.
- Opmeer, E.M., Kortekaas, R., van Tol,M.J., van der Wee N.J., Woudstra, S., van Buchem, M.A., Penninx, B.W., Veltman, D.J., Aleman, A. (2013). Influence of COMT Val158Met genotype on the depressed brain during emotional processing and working memory. PLoS One, 8, 73290
- Polesskaya, O.O. (2004). Different expression of the "C" and "T" alleles of the 5-HT2A receptor gene in the temporal cortex of normal individuals and schizophrenics. Archive of Neurology, 61, 1249.
- Popova, N.K., Il'chibaeva, T.V., Naumenko, V.S. (2017). Nejrotroficheskie faktory (BDNF, GDNF) i serotoninergicheskaja sistema mozga. Obzor [Neurotrophic factors (BDNF, GDNF) and serotoninergic brain system: a review]. Biohimija, 82, 449-459 (in Russian).
- Sergienko, E.A., Vetrova, I.I. (2010). Test D. Mejera, P. Seloveja i D. Karuzo «Emocional'nyj intellekt» (MSCEIT v. 2.0) [Test of D. Mayer, P Salovey, and D. Caruso “Emotional Intelligence” (MSCEIT v. 2.0)]. Moscow: Izd-vo Instituta psihologii RAN (in Russian).
- Tautkute, I., Trzcinski, T., & Bielski, A. (2018). I Know How You Feel: Emotion Recognition with Facial Landmarks. arXiv:1805.00326.
- Wang, L., Ashley-Koch, A., Steffens, D.C., Krishnan, K.R., Taylor, W.D. (2012). Impact of BDNF Val66Met and 5-HTTLPR polymorphism variants on neural substrates related to sadness and executive function. Genes, Brain and Behavior, 11, 352-359.
- Wasser, C.I., Evans, F., Kempnich, C., Glikmann-Johnston, Y., Andrews, S. C., Thyagarajan, D., & Stout, J. C. (2018). Emotion recognition in Parkinson’s disease: Static and dynamic factors. Neuropsychology, 32, 230-234.
- Wingenbach, T.S.H., Ashwin, C., Brosnan, M. (2018) Sex differences in facial emotion recognition across varying expression intensity levels from videos. PLoS One, 13, e0190634.
- Woudstra, S., Bochdanovits, Z., van Tol, M.J., Veltman, D.J., Zitman, F.G., van Buchem, M.A., van der Wee, N.J., Opmeer, E.M., Demenescu, L.R., Aleman, A., Penninx, B.W., Hoogendijk, W.J. (2012) Piccolo genotype modulates neural correlates of emotion processing but not executive functioning. Translational Psychiatry, 2, e99.
- Wright, R., Riedel, R., Sechrest, L., Lane, R. D., & Smith, R. (2018). Sex differences in emotion recognition ability: The mediating role of trait emotional awareness. Motivation and Emotion, 42, 149-160.
Copyright information

This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License.
About this article
Publication Date
23 November 2018
Article Doi
eBook ISBN
978-1-80296-048-8
Publisher
Future Academy
Volume
49
Print ISBN (optional)
-
Edition Number
1st Edition
Pages
1-840
Subjects
Educational psychology, child psychology, developmental psychology, cognitive psychology
Cite this article as:
Kosonogov, V., Vorobyeva, E., Kovsh, E., Titova, A., & Ermakov, P. N. (2018). Association Of Neurotransmitter System Genes With Facial Emotion Recognition And Emotional Intelligence. In S. Malykh, & E. Nikulchev (Eds.), Psychology and Education - ICPE 2018, vol 49. European Proceedings of Social and Behavioural Sciences (pp. 294-301). Future Academy. https://doi.org/10.15405/epsbs.2018.11.02.33

